Article Text
Abstract
Objective Gastric cancer is the second leading cause of cancer-related deaths and the fifth most common malignancy worldwide. In this study, human and mouse gastric cancer organoids were generated to model the disease and perform drug testing to delineate treatment strategies.
Design Human gastric cancer organoid cultures were established, samples classified according to their molecular profile and their response to conventional chemotherapeutics tested. Targeted treatment was performed according to specific druggable mutations. Mouse gastric cancer organoid cultures were generated carrying molecular subtype-specific alterations.
Results Twenty human gastric cancer organoid cultures were established and four selected for a comprehensive in-depth analysis. Organoids demonstrated divergent growth characteristics and morphologies. Immunohistochemistry showed similar characteristics to the corresponding primary tissue. A divergent response to 5-fluoruracil, oxaliplatin, irinotecan, epirubicin and docetaxel treatment was observed. Whole genome sequencing revealed a mutational spectrum that corresponded to the previously identified microsatellite instable, genomic stable and chromosomal instable subtypes of gastric cancer. The mutational landscape allowed targeted therapy with trastuzumab for ERBB2 alterations and palbociclib for CDKN2A loss. Mouse cancer organoids carrying Kras and Tp53 or Apc and Cdh1 mutations were characterised and serve as model system to study the signalling of induced pathways.
Conclusion We generated human and mouse gastric cancer organoids modelling typical characteristics and altered pathways of human gastric cancer. Successful interference with activated pathways demonstrates their potential usefulness as living biomarkers for therapy response testing.
- gastric cancer
This is an open access article distributed in accordance with the Creative Commons Attribution Non Commercial (CC BY-NC 4.0) license, which permits others to distribute, remix, adapt, build upon this work non-commercially, and license their derivative works on different terms, provided the original work is properly cited and the use is non-commercial. See: http://creativecommons.org/licenses/by-nc/4.0/
Statistics from Altmetric.com
Significance of this study
What is already known on this subject?
Organoids recapitulate many aspects of the organ they are derived from.
Cancer organoids have proven to constitute a valuable tool in between 2D cell line-based drug screens and patient trials.
What are the new findings?
Human gastric cancer biobanks can be established.
Human gastric cancer organoids show divergent responses to chemotherapy.
Targeted therapy testing is feasible using gastric cancer organoids.
Mouse gastric cancer organoids serve as model systems with a defined genetic background.
How might it impact on clinical practice in the foreseeable future?
Gastric cancer organoids might serve as living biomarkers to predict therapy response and resistance in individual patients, thereby guiding personalised therapy approaches.
Introduction
Gastric cancer ranks as the fifth most common malignancy and the second leading cause of cancer-related deaths worldwide.1 In addition, adenocarcinomas of the oesophagogastric junction (AEG) constitute an entity with rising incidence rates and histological overlap to gastric cancer.2 3 Several histological subtypes of gastric cancer can be differentiated of which more than 90% are adenocarcinomas. While the WHO classifies four different main types of gastric adenocarcinoma,4 the Lauren’s criteria as the most widely used classification differentiate into an intestinal, diffuse and mixed type.5 The Cancer Genome Atlas (TCGA) Research Network characterised gastric adenocarcinoma into four different molecular subgroups6: those positive for the Epstein-Barr virus (EBV) with frequent PIK3CA mutations and CDKN2A silencing, a microsatellite instable (MSI) subtype with a hypermutation phenotype, a genomically stable (GS) subtype displaying diffuse histology and frequent CDH1 and RHOA mutations and a chromosomal instable (CIN) subtype displaying aneuploidy and frequent mutation of TP53 as well as activation of the receptor tyrosine kinase (RTK)-RAS pathway. Molecular characterisation of AEG revealed their high similarity to the CIN subtype of gastric cancer.7
The prognosis of gastric cancer is often poor. Frequently, the lack of clinical signs leads to a delayed diagnosis with three quarters of patients presenting with non-curable advanced disease.8 Surgery is the only curative option. In addition, interdisciplinary approaches including neoadjuvant and adjuvant chemotherapy have led to improved survival rates.9 10 The most commonly used chemotherapeutic drugs for gastric cancer are fluoropyrimidines (ie, 5-fluoruracil (5-FU), capecitabine, S-1), platinum compounds (ie, cisplatin, oxaliplatin), docetaxel and epirubicin.9–11 Besides classical chemotherapy, genetic alterations represent molecular targets for novel treatment options. So far, the only approved targeted therapies are trastuzumab, a monoclonal antibody inhibiting the epidermal growth factor receptor (HER)-2 signalling and the anti-vascular endothelial growth factor (VEGF) antibody ramucirumab.12 13 Other targeted therapeutics such as the anti-epidermal growth factor receptor (EGFR) antibodies cetuximab and panitumumab or the anti-VEGF antibody bevacizumab have failed to improve survival rates. One reason might be the missing availability of relevant biomarkers to direct targeted therapies to the right patients.
A recently developed three-dimensional (3D) culture system termed ‘organoids’ opens up new opportunities in preclinical personalised therapy testing. Initially developed based on the growth requirements of small intestinal stem cells, organoids have by now been developed for several organs including gastric pyloric and corpus stem cells.14–17 Organoids faithfully recapitulate many aspects of the tissue they are derived from such as differentiation capacity to the tissue-specific lineages as well as stem cell self-renewal.18 19 Human gastric organoids have been shown to constitute a valuable tool to study pathogen infections.20 21 Based on the culture methods for normal tissue successful protocols have been developed for several human cancers.22–24 Advantages of the organoid cultures are the short time frame to establishment compared with xenograft models and the ease of manipulation.25 26 Large organoid collections of individual patient samples function as living human biobanks. Their usefulness has been demonstrated, for example, for a colorectal cancer biobank that was amenable to drug screens to individualise patient treatment and screen for novel therapeutics.27 In addition to primary cancer-derived organoids, establishment of GI organoids from metastatic lesions is feasible and treatment of these organoids recapitulates the clinical response of the corresponding patients.28
In this study, we refined the culture protocol for human gastric cancer organoids, classified the samples according to their molecular profile, assessed their chemotherapeutic response and performed targeted treatments according to specific druggable mutations.
Materials and methods
Human tissues
Human gastric cancer and normal gastric tissues were obtained from patients who underwent surgery at the Department of Visceral, Thoracic and Vascular Surgery at the University Hospital Carl Gustav Carus of the TU Dresden. Clinical data including information on neoadjuvant treatment are summarised in online supplementary table 1.
Supplemental material
Human organoid culture
Human gastric cancer and normal gastric organoids were cultured as described earlier and were passaged twice a week with a split ratio of 1:2/1:3.20 Treatment with chemotherapeutics was performed 24 hours after seeding using 5-FU, oxaliplatin, irinotecan, epirubicin and docetaxel. Selected organoids were treated with trastuzumab (Herceptin, Roche), palbociclib (No S1116, Selleckchem) and imatinib (No ST1571, Selleckchem). Further information is given in the online supplementary methods.
Supplemental material
Whole genome sequencing and RNA sequencing
Whole genome DNA sequencing of organoids was performed with the HighSegXten (Illumina) using a total of 340 ng DNA. DNA was isolated via phenol/chloroform extraction and isopropanol precipitation according to standard protocol. Organoid RNA was isolated using the RNeasy Mini Kit (Qiagen). Sequencing was performed with the NextSeq500 (Illumina) after enrichment of samples with the TruSeq Stranded RNA Library Prep Kit (Illumina) using a total of 200 ng RNA. Detailed methods for bioinformatic processing can be found in the online supplementary methods.
Pathway analysis
A gene set enrichment analysis was done using the R package fgsea and KEGG pathways and Gene Ontology terms.29 30 −log10 (p value) * log2 (fold change) was used as rank function and 100 000 permutations for p value calculation of pathway enrichments. KEGG pathways were plotted using the R package pathview.31
Statistical analysis
Statistical analyses were performed using Student’s t-test and compared normal versus cancer or cancer treated versus untreated (*<0.05; **<0.01; ***<0.001). Graphs were generated using Prism (GraphPad). IC50 calculation was determined with non-linear regression. Western blot experiments were quantified using ImageJ (https://imagej.nih.gov/ij/).
Results
Human gastric cancer organoid culture
Tissue with histologically diagnosed gastric or oesophagogastric adenocarcinoma was obtained from surgical resection specimens (figure 1A). Cultures were initiated using the same organoid medium as for normal stomach tissue.20 To increase successful initiation of organoid cultures and minimise the advent of carrying along normal gastric glands, the tissue digestion method and time as well as processing of isolated tumour patches was optimised (described in the online supplementary methods). Overall, 20 gastric cancer organoid cultures could be initiated. We selected four organoid lines for further analyses based on their high growth rate with a splitting ratio of 1:2/1:3 twice a week. These organoid lines showed divergent morphological phenotypes, indicating different molecular genotypes (figure 1B). DD107 (originating from a corpus carcinoma) had a cystic structure with a thickened epithelium. DD109 (AEG I) demonstrated a non-coherent grape-like growth pattern. DD191 (AEG II) and DD282 (antrum carcinoma) had a compact morphology with no lumen, with DD282 being extremely resistant to dissociation. Normal gastric organoids (DD320N, DD379N, DD392N, DD399N) present a single layered epithelium and cyst-like structure. All organoid lines were continuously in culture for over 1 year without a change in growth behaviour or morphological phenotype. Also, freeze and thaw cycles did not affect or change these growth characteristics (data not shown). Gastric cancer organoids proliferated at different rates, but all with a higher percentage than normal organoids (figure 1C). In order to characterise growth factor dependency, we omitted one by one each relevant media compound for normal organoid growth (figure 1D). Overall, response differed markedly between organoid lines. Normal organoids did grow long term only in complete medium. Omission of A38-01, Fgf10 and Wnt was without phenotypical impact on the cancer lines. Noggin, Egf and the combination of Wnt plus Rspondin were important to varying degrees. Plating of organoids in 2D in plastic flasks led to an attachment of cells, but further passaging either by mechanical dissociation or enzymatic digestion was not possible (data not shown). In a xenograft transplantation experiment cancer organoid lines were subcutaneously injected on the hind legs of mice. Tumour growth was observed for all organoid lines in each mouse (figure 1E). Human origin of xenografts was proven by antinucleoli staining (online supplementary figure 1). The organoid line showing the lowest in vitro proliferation rate (DD191) also showed the slowest in vivo growth.
Supplemental material
Human gastric cancer organoid cultures expand in vitro. (A) Scheme of cancer organoid culture establishment and performed assays. (B) Representative pictures of four different cancer organoid lines and one normal line (scale bar 100 µm). (C) Proliferation rate of gastric organoids assessed by an EdU proliferation assay. (D) Media component depletion. Each annotated component was omitted from the complete medium and organoids followed over 5 weeks. (E) Growth curve of organoid xenografts (n=3 for each organoid line).
To analyse if organoids maintain similar characteristics in culture as the cancers they were derived from, we performed different immunohistochemical reactions for typical gastric cancer markers, that is, cytokeratin 7, cadherin 17, carcinoembryonal antigen and periodic acid Schiff reaction (figure 2A–D). These reactions were compared with the primary tissue as well as with the xenograft tumours. The organoid lines as well as the xenografts derived thereof faithfully and permanently recapitulated the immunohistochemical characteristics of their corresponding primary cancers.
Human gastric cancer organoids maintain primary tumour characteristics. Immunohistochemistry of (A) DD107, (B) DD109, (C) DD191 and (D) DD282 using HE, carcinoembryonal antigen (CEA), cytokeratin 7 (CK7), cadherin 17 (Cadh17) and periodic acid Schiff (PAS) reaction of primary cancers, organoids and organoid-derived xenograft tumours (scale bar 50 µm).
Divergent response of gastric cancer organoids to conventional chemotherapeutics
Patients with gastric cancer treated in neoadjuvant intention with conventional chemotherapeutics respond to varying degrees, documented by their histological regression grade.32 To investigate if gastric cancer organoids reflect this divergent response we treated the organoid lines with conventional chemotherapeutics routinely used in gastric cancer treatment, that is, 5-FU, oxaliplatin, irinotecan, epirubicin and docetaxel. Cell viability assays documented divergent responses to treatments (figure 3A–E). Especially for 5-FU, irinotecan and epirubicin the response of the four lines varied widely. In line with the viability assay results, responding organoids became dark and started to disaggregate (online supplementary figure 3A–E). The divergent response to 5-FU treatment was further analysed by annexin V/propidium iodide flow cytometry in the two lines showing the greatest difference in their response (example experiment in figure 3F). The less responsive line DD109 in the viability assay showed under normal conditions on average 64.0% living cells, which shifted slightly to 61.7% upon treatment with 1 µM 5-FU. On the other hand, in the responsive line DD282 annexin V staining increased from on average 11.6% to 41.7% for early apoptotic cells and 6.5% to 16.4% for late apoptotic cells upon treatment. Only on average 30.0% of cells were unaffected by the treatment, in line with the cell viability results documenting responsiveness of DD282 to 5-FU. In general, patterns of resistance could be documented, for example, DD109 was relatively resistant to 5-FU and epirubicin, while DD191 responded well to the same drugs. At the same time, DD109 responded well to irinotecan treatment, while DD107 showed a response only at higher concentrations. Normal organoids responded in a similar range to chemotherapeutics as the cancer organoids and displayed especially for 5-FU and oxaliplatin differential responses between the organoid lines (online supplementary figure 2 and table 2). For comparative reasons, chemotherapy response was in addition analysed for several classical 2D gastric cancer cell lines (ie, AGS, KatoIII, Snu1 and Snu5) (online supplementary figure 4). A tendency to higher IC50 values and thus more resistance was observed in organoid lines, especially for oxaliplatin and irinotecan (online supplementary table 2). Taken together, at least for in vitro treatment, an active conventional chemotherapeutic drug could be defined for each cancer line, as well as potential resistance patterns.
Supplemental material
Supplemental material
Supplemental material
Human gastric cancer organoids show divergent therapy response to conventional chemotherapeutics. (A–E) Cell viability assay after treatment with 5-fluoruracil (5-FU), oxaliplatin, irinotecan, epirubicin and docetaxel in varying concentrations. Analysis for oxaliplatin, irinotecan, epirubicin and docetaxel treat was performed after 24 hours. Organoids treated with 5-FU were analysed 72 hours post-treatment. Values were normalised to untreated control organoids of the same patient. (F) Apoptosis assay using annexin V/propidium iodide (PI) flow cytometry (one example of three independent experiments is shown).
Mutational landscape of gastric cancer organoids
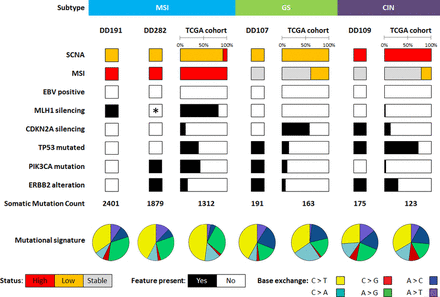
Genomic DNA from organoids and corresponding normal tissue (leucocytes) as well as organoid RNA were subjected to a detailed genomic and transcriptomic analysis. Additionally, the primary tumours were sequenced to verify mutations in the originating tissue. Whole genome sequencing revealed a broad mutational spectrum with features matching DD191 and DD282 to the previously described MSI subtype, DD107 to the GS subtype and DD109 to the CIN subtype (figure 4).6 No genetic EBV traces were detected in organoid lines. The mutation rate per megabasepair (Mb) varied among the samples, showing low mutation rates in DD107 and DD109 (32 and 42 Mut/Mb respectively) as well as a high mutation rate in DD191 (445 Mut/Mb) and DD282 (297 Mut/Mb). A similar pattern was found in overall somatic mutation count with 191 and 175 exonic or splice site mutations in DD107 and DD109, respectively, and 2401 and 1814 exonic or splice site mutations in DD191 and DD282, respectively (online supplementary table 3). These values were comparable to the mean values observed in the TCGA subtypes GS, CIN and MSI with 163, 123 and 1312 mutations, respectively (figure 4).
Supplemental material
Molecular features of gastric cancer organoids in comparison to the The Cancer Genome Atlas (TCGA) cohort. Cancer organoid lines were assigned to the microsatellite instable (MSI), genomic stable (GS) and chromosomal instable (CIN) subtypes. Represented are somatic copy number aberrations (SCNA), microsatellite instability (MSI) status, Epstein-Barr virus (EBV) status, prominent genes and mutational features. Statuses are indicated as high (red), low (orange) and stable (light grey) and presence or absence of features in black or white respectively. *DD282 carries an MSH6 mutation conveying microsatellite instability.
Frequently mutated genes in gastric cancer were detected in our cohort. MLH1 and MSH6 silencing, both critical genes in DNA mismatch repair, associated with microsatellite instability, were present as a Lys435fs mutation in DD191 and an Arg248fs in DD282, respectively. CDKN2A loss, present in most GS cases, was found as a biallelic deletion in DD109 with consequent significant upregulation of CDK4/6-E2F downstream targets BRCA1 and MYB (online supplementary table 3).33 34 TP53, essentially mutated in CIN cases, exhibited a stop Arg280* mutation in DD109. An activating heterozygous Glu542Lys mutation of PIK3CA, also present in TCGA EBV, MSI and GS subtypes, was found in DD107 and three activating heterozygous mutations, namely Arg88Gln, Cys378Arg and Asp805Asn, were found in DD282. A major ERBB2 amplification as well as an activating Ser310Phe mutation was detected in DD109 and DD107, respectively. To evaluate if these two different alterations indeed lead to a RTK-RAS pathway activation, RNASeq data of ERBB2 target genes were analysed. Both the ERBB2 amplification as well as the Ser280Phe mutation effected significantly the c-MYC mediated expression of target genes CCND2, CDKN1A and THBS1 (online supplementary table 3).35 36 Further cancer-associated mutations comprised missense mutations in CDK10 (Gly249Arg), HRAS (Gly13Asp), FGFR2 (Glu201Lys) and SUFU (Asp182Asn) in DD107 (online supplementary table 3 and figure 5). DD109 showed one heterozygous stop mutation in ARID1A (Gln1365*) while DD282 exhibited two heterozygous mutations (Gln1614* and Leu1816fs). The mutational signature showed predominantly C>T and A>G base exchanges in DD191 and DD282, C>T, A>C and to a minor degree A>G base exchanges in DD107, whereas DD109 had an equal distribution of all kinds of base exchanges. A similar distribution was found in the TCGA samples (figure 4).
Circos plots of overall genetic and chromosomal characteristics of gastric cancer organoids. Represented are DD107 (A), DD191 (B), DD109 (C) and DD282 (D). From the outside to the inside: known oncogenes (green: missense, black: non-sense, red: amplifications, blue: deletions), chromosomes, coding variants represented as small squares (green: missense, black: stop or splice site, red: frameshift), copy number variations (CNV) as a scatterplot and as gained (red) or lost (blue) regions as well as interchromosomal (red) and intrachromosomal (blue) rearrangements bigger than 1 Mb. CIN, chromosomal instable; GS, genomically stable; MSI, microsatellite instable.
Somatic copy number aberrations (SCNA) were present in all cases. DD107 showed little chromosomal rearrangements while the genome of DD109 was strongly rearranged (online supplementary figure 5A), thus showing typical features of the GS and CIN subtype, respectively. Especially chromosomes 9, 11 and 17 of DD109 showed strong intrachromosomal rearrangements, while translocations were found between various other chromosomes of its genome. DD191 and DD282 did not have any interchromosomal rearrangement but exhibited high microsatellite instability (figure 5C,D), which was absent in the other two cases.
Supplemental material
Taken together, DD191 and DD282 were assigned to the MSI subtype due to the presence of MLH1 and MSH6 mutations and the microsatellite instability. Genomic stability, PIK3CA as well as ERBB2 mutations matched DD107 to the GS subtype. The highly rearranged genome of DD109 plus the TP53 mutation and the ERBB2 amplification were considered features matching this organoid line to the CIN subtype. All organoid lines were cultured with the same culture medium indicating that the culture method does not exclude certain molecular subtypes. Sequencing of the primary tumours demonstrated conservation of driver mutations and copy number aberrations in the organoids (online supplementary figure 5B,C and table 3).
Targeted therapy testing in gastric cancer organoids
Personalised anticancer treatment holds great promises for the future of oncology.37 Nevertheless, many approaches to target specific signalling pathways in gastric cancer including the EGFR, hepatocyte growth factor/MET or fibroblast growth factor receptor (FGFR) pathway have failed; one reason is the lack of predictive biomarkers for therapy response.38 To explore the potential use of gastric organoids as living biomarkers for therapy response, activated pathways based on the mutation pattern of individual organoid lines were defined (online supplementary table 4). We observed a known activating HER2 receptor mutation (Ser310Phe) in DD107 and a variant of unknown significance (VUS; Gly201Asp) in DD282. The mutation Ser310Phe is located in the extracellular domain of the receptor and is drug sensitive.6 39 In addition, we found an amplification of the ERBB2 receptor in DD109, which could be confirmed by immunohistochemistry (figure 6A). Anti-HER2 treatment using the antibody trastuzumab resulted in a viability decrease to 70.6% for DD107, to 79.2% for DD109 and to 85.6% for DD282 (figure 6B). The VUS in DD282 might therefore indeed constitute an activating mutation. In order to explore the feasibility of testing combinatorial treatment regiments, the two lines carrying known Her2 receptor activating alterations (DD107 and DD109) were treated with trastuzumab plus 5-FU.13 An additive effect of trastuzumab compared with 5-FU alone could be observed in both organoid lines with complete death at the highest concentration, whereas 5-FU alone only could lead to a reduction of viability to a maximum of 40% (online supplementary figure 6). To further characterise the effect of trastuzumab treatment we analysed the phosphorylation level of the extracellular-signal regulated kinase 1/2 (ERK1/2), a downstream member of the RAS/RAF signalling cascade. While for DD107 and DD282 no change was detected, DD109 downregulated the phosphorylation level of ERK1/2 by 55% (figure 6C,D). Apparently, DD107, which showed the highest response upon trastuzumab treatment and carries an activating Her2 mutation, signals through a RAS/RAF independent pathway, while DD109 with a classical amplification downregulates the RAS/RAF signalling cascade.
Supplemental material
Supplemental material
Targeting of mutated pathways in human gastric cancer organoids. (A) Immunohistochemistry for ERBB2 confirming the genomic amplification in DD109 (scale bar 50 µm). (B) Cell viability measurement after 72 hours of 0.1 µM trastuzumab. Student’s t-test treated versus untreated; **<0.01; ***<0.001. Western blot (C) and densitometric analysis (D) for ERK1/2 and phospho-ERK1/2 after 72 hours of 0.1 µM trastuzumab. (E) Western blot for phospho-c-KIT 24 hours after 100 µM imatinib treatment on DD109 (p.Glu142Asp c-KIT mutation). (F) Targeting of CDK4/6 in DD109 (biallelic loss of CDKN2A) and DD191 (no CDKN2A alteration) as well as DD320N (normal control) using 5 µM palbociclib. Proliferation was analysed after 24 hours of treatment using an EdU assay. (G) Long-term treatment (11 days, two splits) using 10 µM palbociclib led to the loss of the DD191 and DD320N culture (scale bar 100 µm).
The DD109 organoid line carried a mutation in the stem cell growth factor receptor (c-KIT) gene. The mutation was located in exon 3 (p.Glu142Asp), which encodes part of the extracellular domain of the receptor. A mutation in this amino acid of c-KIT has to our knowledge not been described before; the effect of this mutation on c-KIT signalling is therefore unknown. Treatment with 100 µM imatinib, a tyrosine kinase inhibitor that blocks activation of c-KIT, resulted in a dephosphorylation of the receptor (figure 6E).
Additionally, DD109 harboured a biallelic DNA loss in the CDKN2A locus. CDKN2A encodes for the tumour suppressor p16, which plays an important role in regulating the cell cycle via inhibiting the cyclin-dependent kinase 4/6 (CDK4/6). We treated DD109, DD191 (no mutation in CDKN2A) and DD320N (normal control) organoids with palbociclib, a potent inhibitor of CDK4/6. A complete block in proliferation was detected for DD191 as well as the normal DD320N in an EdU proliferation assay, while DD109 showed a strong decrease with 2% of cells still cycling (figure 6F). Long-term treatment with palbociclib including two times passaging resulted in a loss of the DD191 and DD320N cultures, indicating a complete inhibition of proliferation. DD109 organoids were still present and phenotypically unharmed (figure 6G), while their growth measured by split rate as well as in the EdU assay was inhibited. We concluded that the loss of p16 in DD109 led to the survival of the line due to insufficient blockage of CDK4/6 by palbociclib.
Mouse organoids allow gastric cancer modelling with defined mutation spectrum
Human gastric cancer organoids allow a detailed analysis of interference into an activated pathway for an individual patient. Due to the high total number of mutations (on average 140 in the GS and CIN subtypes) effects of targeted drugs always need to be interpreted in the context of the complete mutation spectrum present in the specific organoid line. In order to set up models for in-depth analyses of pathway interference in a defined mutation spectrum, we chose to combine inducible alleles frequently mutated in the CIN and GS subtype in a mouse model. The CIN subtype is characterised by TP53 mutations and active RTK-RAS signalling. Based on own calculations of publicly available data from the TCGA data set,6 the mutations are present in 80% and 57% of cases, respectively, and in combination affect 48% of CIN cases. In order to model the CIN subtype of gastric cancer, inducible mutated alleles of the two most characteristic mutations, that is, KrasG12D and Tp53R172H , were combined (CIN model).6 40 41 The GS subtype carries mutations in the cell motility genes CDH1 and RHOA in 45% of cases. To model the GS subtype a floxed allele of Cdh1 was used. In addition, GS cancers frequently harbour oncogenic driver mutations in the transforming growth factor beta (TGF-β) (33%), the RTK/RAS (31%) and the WNT pathway (24%).6 We chose to model an activated Wnt pathway using a floxed Apc allele (GS-WNT model).42 As a stomach-specific induction in vivo is currently not possible due to the absence of a stomach-specific Cre mouse line, the inducible alleles were recombined in vitro. After establishing normal gastric organoids from mouse gastric corpus, recombination was induced using a Cre-GFP expressing adenovirus and confirmed by genotyping (online supplementary figure 7).
Supplemental material
Tumour modelling using corpus mouse organoids simulating the chromosomal instable (CIN) and genomic stable (GS) subtype of stomach cancer. (A–H) Established tumour organoids were analysed concerning mutated pathways and compared with normal organoids. (A–B, E–F) Representative pictures of organoids showed differences in morphology. CIN tumours showed smooth layered epithelium compared with normal organoids. GS-WNT tumour organoids had a grape-like structure. (C, G) Immunohistochemistry of normal versus tumour organoids. CIN organoids showed a Tp53 accumulation in the nucleus. GS-WNT organoids showed β-catenin accumulation in the nucleus whereas normal organoids showed membranous expression. (D) Kras pathway of CIN organoids was quantified by Western blot. Tumour organoids were increased in Erk1/2 phosphorylation compared with normal organoids. (H) Quantitative RT-PCR experiments of GS-Wnt organoids. Tumours showed an increased Wnt signalling as measured by Axin2 and Cyclin D1 expression levels compared with normal organoids. Treatment with 5 mM calphostin C decreased pathway activity (scale bar A–G 100 µm, Student’s t-test treated vs non-treated *<0.05; ** < 0.01). (I) Media component release. Each indicated medium component was omitted from the whole medium one by one and organoids passaged two times per week in a 1:2 ratio. (J) Single cell plating of organoids. One hundred single cells were plated per well and originating organoids counted after 7 days. Student’s t-test tumour versus control (*<0.05).
The CIN model showed a thickened irregular epithelium, which was partly multilayered, thus phenotypically differing from the homogenous single-layered normal gastric organoids (figure 7A,B). The Tp53 mutation resulted in a nuclear accumulation of the corresponding protein (figure 7C). Activated Rtk/Ras signalling was documented by a strong Erk1/2 phosphorylation (figure 7D). The model therefore recapitulates key features of CIN cancers: Tp53 mutation and Rtk/Ras pathway activation.
The Cdh1 mutation in the GS-WNT model resulted in a complete structural alteration towards a grape-like form compared with normal organoids, induced by the loss of Cdh1/E-cadherin-mediated cell-cell connections (figure 7E,F). The Apc loss resulted in aberrant Wnt signalling observable by nuclear accumulation of beta-Catenin (Ctnnb1) and increased expression of the Wnt signature genes Axin2 and CyclinD1 (Ccnd1) (figure 7G,H). Treatment of the GS-WNT model with calphostin C, a potent inhibitor of the β-catenin/TCF complex, reduced downstream Wnt target gene expression (figure 7H). The GS-WNT organoid line therefore constitutes a model system to test the effect of targeted drugs for Wnt-activated GS cancers.
To further characterise the transformation of organoids after viral infection we released Noggin, Wnt, Wnt plus Rspondin, Fgf10 and Egf from the media and compared the CIN and GS-WNT model with normal organoids (figure 7I). Tumour organoids were unaffected whereas the normal organoid line was lost at different time points after growth factor withdrawal. To test the ability of the different lines to initiate organoids from single cells we plated 100 single cells per well and counted the formed organoids after 7-day culturing (figure 7J). An increase in organoid formation efficiency could be documented for the tumour lines, especially in the GS-WNT model. Similar to the human organoid lines, it was not possible to culture the mouse tumour organoid lines in plastic flasks over longer time span (data not shown). Taken together, alterations in only two cancer genes per model resulted in a robust transformation of the organoid lines modelling molecular subtypes of gastric cancer.
Discussion
Here we demonstrate the usefulness of mouse and human gastric cancer organoids to study molecular mutation patterns and their effect on drug sensitivity. Mouse cancer organoids with defined genetic alterations circumvent the classical problem of working with cancer cell lines with a complex mutational landscape difficult to interpret. These models allow the detailed analysis of interference with individual pathways in a defined setting. Human organoid lines derived from primary cancers, on the other hand, are models to test the effect of a certain targeted drug for an individual patient. Using bioinformatics, predictions can be made on the potential effectiveness of a given drug for a specific mutation, but often the highly altered genetic background in an individual patient with cancer hampers precise predictions, as different activated or deactivated pathways can interfere with each other. Human cancer organoid lines allow drug testing in a living system, with the response or resistance being the result of the concurrence of all present mutations of an individual patient. It has to be kept in mind that organoids consist only of the epithelial layer with no surrounding mesenchyme, blood vessels or immune cells. Drugs that target the microenvironment of a cancer thus cannot be evaluated.
In this study, we illustrate the possibility to establish larger organoid biobanks also for gastric cancer, similar to what has been shown for colorectal cancer, prostate cancer and pancreatic cancer.22 24 27 Focusing on four gastric cancer organoid lines with divergent phenotypes concerning their growth pattern (figure 1), we first analysed their phenotype using immunohistochemical staining typically performed to pathologically characterise gastric cancer. The organoids as well as their xenografts hereby phenocopied the architecture of the primary cancers they were derived from (figure 2). Next, we set out to analyse the response of these organoid lines to classical chemotherapy. Employed drugs were selected based on currently used clinical treatment schemes.9–11 Organoids were exposed to 5-FU, oxaliplatin, irinotecan, docetaxel and epirubicin, and for each drug divergent responses could be documented (figure 3). As an example, DD109 showed a response to 5-FU only at a high concentration, while DD191 and DD282 responded already at low doses (figure 3A). For each organoid line an active chemotherapeutic drug could be defined: for example, DD109 showed resistance to most applied drugs except irinotecan (figure 3C). Just as important for an individual patient as to define a ‘working’ drug is the definition of drugs for which an inherent resistance is present, thus most likely causing only side effects and little therapeutic response.
Whole genome sequencing allowed us to match the analysed organoid lines to recently described molecular subtypes of gastric cancer,6 that is, DD107 matched to the GS, DD109 to the CIN and DD191 as well as DD282 to the MSI subtype, taking into consideration somatic mutations and structural variations including SCNAs (figures 4 and 5). RNASeq data provided additional insights into downstream target gene and pathway activations. The identification of mutations in known oncogenes or tumour suppressor genes allowed us to test targeted therapy approaches in our organoid lines (online supplementary table 4). DD107 carried a known activating mutation and DD282 an unknown variant, whereas DD109 showed a gene amplification in the ERBB2 gene (HER2/neu). Amplifications of ERBB2 are found in gastric cancer in up to 22% of cases and can be successfully targeted with trastuzumab, an antibody binding to the HER2 receptor, showing a significant overall survival benefit in the clinical setting.13 Interestingly, we found a response upon trastuzumab treatment in both the amplified as well as the mutated ERBB2 organoid line (figure 6A). In addition, combining trastuzumab with 5-FU treatment resulted in an additive treatment effect (online supplementary figure 6). The mitogen-activated protein kinase (MAPK) pathway as measured by phosphorylation of ERK1/2 was only downregulated in the amplified ERBB2 organoid line DD109 (figure 6C,D). The mechanism of action on the mutated lines remains unclear as ERBB2 signals through a diverse set of downstream signalling pathways. Treatment of cell lines with ERBB2 mutations was previously shown to result in a mixed response, with some mutations being responsive to trastuzumab, while others were not.39 Using organoids, the effectiveness of a targeted therapy on mutations can thus be tested specifically for each patient.
CDKN2A encodes for the important tumour suppressor p16 playing a key role in cell cycle control and is frequently mutated in gastric cancer.6 DD109 contains a biallelic loss of CDKN2A and we treated the organoid line with palbociclib, a Food and Drug Administration-approved CDK4/6 inhibitor. Organoid growth was arrested in two control lines with functional CDKN2A leading to death, while DD109 continued to show a minimal proliferation in 2% of cells, which was sufficient to keep the line alive (figure 6F,G). A potential explanation is the dual inhibition of the cell cycle by palbociclib and a functional CDK4/6 in the control lines, while in DD109 the loss of CDKN2A resulted in an incomplete suppression of the cell cycle.43
The line DD109 carried an unknown mutation in exon 3 of the c-KIT receptor. Other known mutations also affecting the extracellular domain activate the downstream signalling pathway. This assumption was confirmed by a strong phosphorylation of Tyr719 of c-KIT, indicating active signalling (figure 6E). Treatment with imatinib resulted in a dephosphorylation of the receptor. Organoids can therefore also be used to delineate the effect of treatments on unknown mutations.
As discussed above, the analysis of interference into a specific signalling pathway can be affected by crosstalk from other aberrantly activated signalling pathways. To allow unbiased testing of therapeutic interventions, we generated two mouse organoid models for the CIN and GS molecular subtype with a defined mutational load. Organoids carrying the hotspot mutation Tp53R172H that leads to an accumulation of the truncated p53 protein in the nucleus as well as the hotspot mutation KrasG12D activating the MAPK signalling pathway were generated modelling the CIN subtype. Activating the mutations resulted in a phenotypic change of the organoids towards a multilayered atypical epithelium with increased Erk1/2 phosphorylation indicating active MAPK signalling (figure 7D). Similar observations were also made by Li et al using a collagen-based air-liquid model of neonatal mouse cells directly transformed by adenovirus Cre upon initiation of culture.44 Organoids modelling the GS-WNT subtype showed a grape-like structure leading to a diffuse morphology due to the loss of the cell-cell connections, resembling also the classical diffuse type of gastric cancer according to the Lauren classification. The concomitant loss of Apc led to an increased expression of Wnt target genes, which could be partly reversed by calphostin C treatment (figure 7H). Both tumour models can serve in the future to test therapeutic interventions in a defined genetic background.
In summary, we generated human and mouse cancer organoids representing typical characteristics and altered pathways of human gastric cancer. Successful interference with activated pathways demonstrates their potential usefulness in testing therapy response and resistance for individual patients. For a small number of cases, comparisons of patient to organoid responses have already demonstrated the high predictive power of organoids.28 Including organoids into clinical trials and treating them side by side with the patients from which the organoids were derived will ultimately test their usefulness as a biomarker to predict individual therapy response. Besides therapy response testing, organoids already fill the gap between traditional 2D cell-line based drug screening and clinical trials.
Supplemental material
Supplemental material
Acknowledgments
We thank Susanne Schindler, Alexander Krüger and Katja Schumann for excellent technical assistance as well as the Genomics and Proteomics Core Facility, German Cancer Research Center (DKFZ) for providing excellent high throughput sequencing services.
References
Footnotes
TS, SRM and AR contributed equally.
Contributors Study concept and design: TS, SRM, DES. Acquisition of data: TS, SRM, FZ, CvN, AR, KG, CS, SS, KW, HU, US, DEA, BK. Analysis and interpretation of data: all coauthors. Drafting of the manuscript: TS, SRM, DES. Critical revision of the manuscript for important intellectual content: GBB, TW, DEA, MK, JW. Statistical analysis: TS, SRM, FZ, KG, BK.
Funding Funding was provided by the Deutsche Krebshilfe (No 111350), the Sander Stiftung (No 2014.104.1), the Hector Stiftung (No M65.2) and the European Union (ERC No 639050) to DES. AR was supported by the Preiss-Daimler Stiftung and Medizinischen Fakultät Carl Gustav Carus TU Dresden. AMG was supported by the Deutsche Krebshilfe (No 70112925).
Competing interests None declared.
Patient consent Obtained.
Ethics approval Ethical Committee of the TU Dresden (No EK451122014).
Provenance and peer review Not commissioned; externally peer reviewed.