Abstract
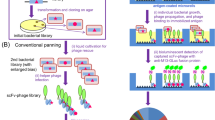
The creation of large phage antibody libraries has become an important goal in selecting antibodies against any antigen. Here we describe a method for making libraries so large that the complete diversity cannot be accessed using traditional phage technology. This involves the creation of a primary phage scFv library in a phagemid vector containing two nonhomologous lox sites. Contrary to the current dogma, we found that infecting Cre recombinase–expressing bacteria by such a primary library at a high multiplicity of infection results in the entry of many different phagemid into the cell. Exchange of Vh and Vl genes between such phagemids creates many new V h/Vl combinations, all of which are functional. On the basis of the observed recombination, the library is calculated to have a diversity of 3×1011. A library created using this method was validated by the selection of high affinity antibodies against a large number of different protein antigens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Barbas, C.F., Kang, A.S., Lerner, R.A. & Benkovic, S.J. Assembly of combinatorial antibody libraries on phage surfaces: the gene III site. Proc. Natl. Acad. Sci. USA 88, 7978– 7982 (1991).
Griffiths, A.D. et al. Human anti-self antibodies with high specificity from phage display libraries. EMBO J. 12, 725– 734 (1993).
Griffiths, A.D. et al. Isolation of high affinity human antibodies directly from large synthetic repertoires EMBO J. 13, 3245–3260 (1994).
Marks, J.D. et al. By-passing immunization: building high affinity human antibodies by chain shuffling. Bio/Technology. 10, 779–783 (1992).
Nissim, A. et al. Antibody fragments from a ‘single pot’ phage display library as immunochemical reagents. EMBO J. 13, 692–698 (1994).
Perelson, A.S. & Oster, G.F. Theoretical studies of clonal selection: minimal antibody repertoire size and reliability of self non-self discrimination . J. Theor. Biol. 81, 645– 670 (1979).
Vaughan, T.J. et al. Human antibodies with sub-nanomolar affinities isolated from a large non-immunized phage display library. Nat. Biotechnol. 14, 309–314 (1996).
de Haard, H.J. et al. A large non-immunized human Fab fragment phage library that permits rapid isolation and kinetic analysis of high affinity antibodies. J. Biol. Chem. 274, 18218–18230 (1999).
Sheets, M.D. et al. Efficient construction of a large nonimmune phage antibody library; the production of panels of high affinity human single-chain antibodies to protein antigens. Proc. Natl. Acad. Sci. USA 95, 6157–6162 (1998).
Geoffroy, F., Sodoyer, R. & Aujame, L. A new phage display system to construct multicombinatorial libraries of very large antibody repertoires. Gene 151, 109–113 (1994).
Tsurushita, N., Fu, H. & Warren, C. Phage display vectors for in vivo recombination if immunoglobulin heavy and light chain genes to make large combinatorial libraries. Gene 172, 59–63 (1996).
Waterhouse, P., Griffiths, A.D., Johnson, K.S. & Winter, G. Combinatorial infection and in vivo recombination: a strategy for making large phage antibody repertoires. Nucleic Acids Res. 21, 2265–2266. (1993).
Hoess, R., Wierzbicki, A. & Abremiski, K. The role of the loxP spacer region in P1 site-specific recombination. Nucleic Acids Res. 14, 2287 –2300 (1986).
Mariuzza, R.A. et al. Preliminary crystallographic study of the complex between the Fab fragment of a monoclonal anti-lysozyme antibody and its antigen. J. Mol. Biol. 170, 1055–1058 (1983).
Furth, M.E., Davis, L.J., Fleurdelys, B. & Scolnic, E.M. Monoclonal antibodies to the p21 products of the transforming gene of Harvey murine sarcoma virus and of the cellular ras gene family. J. Virol. 43, 294–304 ( 1982).
Holliger, P., Prospero, T. & Winter, G. “Diabodies”: small bivalent and bispecific antibody fragments. Proc. Natl. Acad. Sci. USA. 90, 6444–6448 (1993).
Tomlinson, I.M., Williams, S.C., Corbett, S.J., Cox, J.P.L. & Winter, G. VBASE sequence directory.(MRC Centre for Protein Engineering, Cambridge, UK; 1996) http://www.mrc-cpe.cam.ac.uk/imt-doc/vbase-home-page.html.
de Kruif, J., Boel, E. & Logtenberg, T. Selection and application of human single chain Fv antibody fragments from a semi-synthetic phage antibody display library with designed CDR3 regions. J. Mol. Biol. 248, 97– 105 (1995).
Marks, J.D. et al. By-passing immunization. Human antibodies from V-gene libraries displayed on phage. J. Mol. Biol. 222, 581 –597 (1991).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular cloning: a laboratory manual(Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY; 1989).
de Bruin, R., Spelt, K., Mol, J., Koes, R. & Quattrocchio, F. Selection of high-affinity phage antibodies from phage display libraries Nat. Biotechnol.. 17, 397–399 (1999).
Sblattero, D. & Bradbury, A. A definitive set of oligonucleotide primers for amplifying human V regions. Immunotechnology 3, 271–278 (1998).
Schier, R. & Marks, J.D. Efficient in vitro affinity maturation of phage antibodies using BIAcore guided selections. Hum. Antib. Hybrid. 7, 97–105. ( 1996).
Clackson, T. & Wells, J.A. In vitro selection from protein and peptide libraries. Trends Biotechnol. 12, 173–184. (1994).
Schier, R. et al. Isolation of high-affinity monomeric human anti-c-erbB-2 single chain Fv using affinity-driven selection. J. Mol. Biol. 255, 28–43 (1996).
Sauer, B. & Henderson, N. The cyclization of linear DNA in Escherichia coli by site-specific recombination. Gene . 70, 331–341 ( 1988).
Hanke, T., Szawlowski, P. & Randall, R.E. Construction of solid matrix-antibody-antigen complexes containing simian immunodeficiency virus p27 using tag-specific monoclonal antibody and tag-linked antigen. J. Gen. Virol. 73, 653–660 (1992).
Werge, T.M., Bradbury, A., Di Luzio, A. & Cattaneo, A. A recombinant cell line, expressing a form of the anti-p21ras antibody, Y13-259, which binds protein A and may be produced as ascites. Oncogene 7, 1033–1035 ( 1992).
Krebber, A. et al . Reliable cloning of functional antibody variable domains from hybridomas and spleen cell repertoires employing a reengineered phage display system. J. Immunol. Methods 201, 35–55 (1997).
Acknowledgements
We are grateful to Roberto Marzari, Jianlong Lou, and Chonglin Yang for helpful discussions; to Francesco Tedesco for RNA from human lymphocytes; to Gabriella Rossi and Jessica Franzot for excellent technical help; and to the following for the proteins they kindly provided for selection: Min Park (Rad52), Scott Peterson (cyclin D, cdk2, cdc25A), Tom Peat (phosphoglycerate dehydrogenase). We would also like to thank Brian Sauer for providing BS1365 as well as many useful discussions on Cre recombinase. This work was partly financed by Fondo Trieste.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sblattero, D., Bradbury, A. Exploiting recombination in single bacteria to make large phage antibody libraries. Nat Biotechnol 18, 75–80 (2000). https://doi.org/10.1038/71958
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/71958
This article is cited by
-
CRISPR-Cas9 mediated phage therapy as an alternative to antibiotics
Animal Diseases (2023)
-
Insights into next generation sequencing guided antibody selection strategies
Scientific Reports (2023)
-
A pandemic-enabled comparison of discovery platforms demonstrates a naïve antibody library can match the best immune-sourced antibodies
Nature Communications (2022)
-
A single donor is sufficient to produce a highly functional in vitro antibody library
Communications Biology (2021)