Abstract
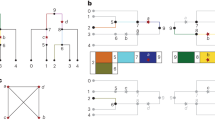
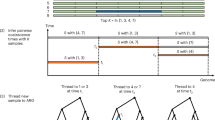
Genome-wide linkage disequilibrium (LD) mapping of common disease genes could be more powerful than linkage analysis if the appropriate density of polymorphic markers were known and if the genotyping effort and cost of producing such an LD map could be reduced. Although different metrics that measure the extent of LD have been evaluated1,2,3, even the most recent studies2,4 have not placed significant emphasis on the most informative and cost-effective method of LD mapping—that based on haplotypes. We have scanned 135 kb of DNA from nine genes, genotyped 122 single-nucleotide polymorphisms (SNPs; approximately 184,000 genotypes) and determined the common haplotypes in a minimum of 384 European individuals for each gene. Here we show how knowledge of the common haplotypes and the SNPs that tag them can be used to (i) explain the often complex patterns of LD between adjacent markers, (ii) reduce genotyping significantly (in this case from 122 to 34 SNPs), (iii) scan the common variation of a gene sensitively and comprehensively and (iv) provide key fine-mapping data within regions of strong LD. Our results also indicate that, at least for the genes studied here, the current version of dbSNP would have been of limited utility for LD mapping because many common haplotypes could not be defined. A directed re-sequencing effort of the approximately 10% of the genome in or near genes in the major ethnic groups would aid the systematic evaluation of the common variant model of common disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Devlin, B. & Risch, N. A comparison of linkage disequilibrium measures for fine-scale mapping. Genomics 29, 311–322 (1995).
Morton, N.E. et al. The optimal measure of allelic association. Proc. Natl Acad. Sci. USA 98, 5217–5221 (2001).
Pritchard, J.K. & Przeworski, M. Linkage disequilibrium in humans: models and data. Am. J. Hum. Genet. 69, 1–14 (2001).
Reich, D.E. et al. Linkage disequilibrium in the human genome. Nature 411, 199–204 (2001).
Daly, M.J., Rioux, J.D., Schaffner, S.F., Hudson, T.J. & Lander, E.S. High-resolution haplotype structure in the human genome. Nature Genet. 29, 229–232.
Rioux, J.D. et al. Genetic variation in the 5q31 cytokine gene cluster confers susceptibility to Crohn's disease. Nature Genet. 29, 223–228.
Tishkoff, S.A. et al. Global patterns of linkage disequilibrium at the CD4 locus and modern human origins. Science 271, 1380–1387 (1996).
Kidd, K.K. et al. A global survey of haplotype frequencies and linkage disequilibrium at the DRD2 locus. Hum. Genet. 103, 211–227 (1998).
Clark, A.G. et al. Haplotype structure and population genetic inferences from nucleotide-sequence variation in human lipoprotein lipase. Am. J. Hum. Genet. 63, 595–612 (1998).
Bonnen, P.E. et al. Haplotypes at ATM identify coding-sequence variation and indicate a region of extensive linkage disequilibrium. Am. J. Hum. Genet. 67, 1437–1451 (2000).
Hollox, E.J. et al. Lactase haplotype diversity in the Old World. Am. J. Hum. Genet. 68, 160–172 (2001).
Rieder, M.J., Taylor, S.L., Clark, A.G. & Nickerson, D.A. Sequence variation in the human angiotensin converting enzyme. Nature Genet. 22, 59–62 (1999).
Harding, R.M. et al. Evidence for variable selective pressures at MC1R. Am. J. Hum. Genet. 66, 1351–1361 (2000).
Fullerton, S.M. et al. Apolipoprotein E variation at the sequence haplotype level: implications for the origin and maintenance of a major human polymorphism. Am. J. Hum. Genet. 67, 881–900 (2000).
Nickerson, D.A. et al. DNA sequence diversity in a 9.7-kb region of the human lipoprotein lipase gene. Nature Genet. 19, 233–240 (1998).
Templeton, A.R., Weiss, K.M., Nickerson, D.A., Boerwinkle, E. & Sing, C.F. Cladistic structure within the human lipoprotein lipase gene and its implications for phenotypic association studies. Genetics 156, 1259–1275 (2000).
Jeffreys, A.J., Kauppi, L. & Neumann, R. Intensely punctate meiotic recombination in the class II region of the major histocompatibility complex. Nature Genet. 29, 217–222.
Martin, E.R. et al. SNPing away at complex diseases: analysis of single-nucleotide polymorphisms around APOE in Alzheimer disease. Am. J. Hum. Genet. 67, 383–394 (2000).
Lucassen, A.M. et al. Susceptibility to insulin dependent diabetes mellitus maps to a 4.1 kb segment of DNA spanning the insulin gene and associated VNTR. Nature Genet. 4, 305–310 (1993).
Nickerson, D.A. et al. Sequence diversity and large-scale typing of SNPs in the human apolipoprotein E gene. Genome Res. 10, 1532–1545 (2000).
Horikawa, Y. et al. Genetic variation in the gene encoding calpain-10 is associated with type 2 diabetes mellitus. Nature Genet. 26, 163–175 (2000).
Ogura, Y. et al. A frameshift mutation in NOD2 associated with susceptibility to Crohn's disease. Nature 411, 603–606 (2001).
Marth, G. et al. Single-nucleotide polymorphisms in the public domain: how useful are they? Nature Genet. 27, 371–372 (2001).
Hugot, J.P. et al. Association of NOD2 leucine-rich repeat variants with susceptibility to Crohn's disease. Nature 411, 599–603 (2001).
Longmate, J.A. Complexity and power in case-control association studies. Am. J. Hum. 68, 1229–1237 (2001).
Drysdale, C.M. et al. Complex promoter and coding region β2-adrenergic receptor haplotypes alter receptor expression and predict in vivo responsiveness. Proc. Natl Acad. Sci. USA 97, 10483–10488 (2000).
Mummidi, S. et al. Evolution of human and non-human primate CC chemokine receptor 5 gene and mRNA. Potential roles for haplotype and mRNA diversity, differential haplotype-specific transcriptional activity, and altered transcription factor binding to polymorphic nucleotides in the pathogenesis of HIV-1 and simian immunodeficiency virus. J. Biol. Chem. 275, 18946–18961 (2000).
Joosten, P.H., Toepoel, M., Mariman, E.C. & Van Zoelen, E.J. Promoter haplotype combinations of the platelet-derived growth factor α-receptor gene predispose to human neural tube defects. Nature Genet. 27, 215–217 (2001).
Todd, J.A. et al. Identification of susceptibility loci for insulin-dependent diabetes mellitus by trans-racial gene mapping. Nature 338, 587–589 (1989).
Degli-Esposti, M.A. et al. Ancestral haplotypes reveal the role of the central MHC in the immunogenetics of IDDM. Immunogenetics 36, 345–356 (1992).
Farrall, M. et al. Fine-mapping of an ancestral recombination breakpoint in DCP1. Nature Genet. 23, 270–271 (1999).
Mein, C.A. et al. Evaluation of single nucleotide polymorphism typing with invader on PCR amplicons and its automation. Genome Res. 10, 330–343 (2000).
Acknowledgements
We are grateful to L. Smink for advice. This work was funded by the Wellcome Trust, the Juvenile Diabetes Research Foundation International, Diabetes UK and grants from the Finnish Academy (38387, 46558, 52114, and 51225), Novo Nordisk Foundation and Sigrid Juselius Foundation. G.C.L.J. was the recipient of a Diabetes UK PhD Studentship.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Johnson, G., Esposito, L., Barratt, B. et al. Haplotype tagging for the identification of common disease genes. Nat Genet 29, 233–237 (2001). https://doi.org/10.1038/ng1001-233
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng1001-233
This article is cited by
-
Association between SLCO1B1 genetic polymorphisms and bleeding risk in patients treated with edoxaban
Scientific Reports (2023)
-
Effect of functional variant rs11466313 on breast cancer susceptibility and TGFB1 promoter activity
Breast Cancer Research and Treatment (2020)
-
Natural variations of FT family genes in soybean varieties covering a wide range of maturity groups
BMC Genomics (2019)
-
Association analysis of the glutelin synthesis genes GluA and GluB1 in a Japonica rice collection
Molecular Breeding (2017)
-
Association of SNP Haplotypes of HKT Family Genes with Salt Tolerance in Indian Wild Rice Germplasm
Rice (2016)